













中国农业科学 ›› 2026, Vol. 59 ›› Issue (1): 29-40.doi: 10.3864/j.issn.0578-1752.2026.01.003
吴琼1( ), 谢香庭2, 王磊2, 牟勇2, 李进伟1,*(
), 谢香庭2, 王磊2, 牟勇2, 李进伟1,*( )
)
收稿日期:2025-07-23
接受日期:2025-10-02
出版日期:2026-01-01
发布日期:2026-01-07
通信作者:
联系方式:
吴琼,E-mail:wuqiong@dbn.com.cn。
基金资助:
WU Qiong1( ), XIE XiangTing2, WANG Lei2, MOU Yong2, LI JinWei1,*(
), XIE XiangTing2, WANG Lei2, MOU Yong2, LI JinWei1,*( )
)
Received:2025-07-23
Accepted:2025-10-02
Published:2026-01-01
Online:2026-01-07
摘要:
【目的】转基因大豆DBN8205转化体已获批转基因生物安全证书(生产应用),转育品种即将产业化应用,建立DBN8205转化体特异性定性定量检测方法,为转基因生物安全监管和定量标识制度的实施提供依据。【方法】根据DBN8205转化体的分子特征序列,设计转化体特异性引物和探针。通过比较多个引物探针组合的扩增曲线及Ct值,筛选出最佳引物探针组合。分别在实时荧光定量PCR和微滴数字PCR平台上考察DBN8205转化体特异性PCR方法的特异性、检测限、定量限、动力学范围和定量准确性等技术参数。制定标准方法联合验证方案,邀请多家实验室对实时荧光定量PCR方法进行联合验证,对联合验证数据进行统计分析,考察方法的重复性和重现性。【结果】筛选出最佳引物探针组合DBN8205-QF/QR/QP,扩增产物长120 bp,仅特异性识别DBN8205转化体成分,具有良好的扩增特异性。在实时荧光定量PCR平台上,检测限为10 copies,定量限为40 copies,标准曲线的各项技术参数符合标准要求,在40—8.2×104 copies动力学范围内,模板拷贝数和Ct值间具有良好的线性关系,能够对含量低至0.1%的样品进行准确定量。8家实验室联合验证结果表明,DBN8205转化体特异性实时荧光定量PCR方法具有良好的重复性和重现性。在微滴数字PCR平台上,检测限和定量限与实时荧光定量PCR方法相同,分别为10和40 copies,动力学范围为40—8.0×104 copies,能够对低至0.1%的样品进行准确定量,定量结果比qPCR具有更高的精密度。t检验表明,实时荧光定量PCR和二重微滴数字PCR的定量结果具有良好的一致性。【结论】建立的DBN8205转化体特异性定量PCR方法能够对DBN8205转化体进行身份鉴定,可在实时荧光定量PCR和微滴数字PCR平台上对产品中的DBN8205转化体成分进行精准定量分析。
吴琼, 谢香庭, 王磊, 牟勇, 李进伟. 转基因大豆DBN8205转化体特异性定量PCR方法的研发和验证[J]. 中国农业科学, 2026, 59(1): 29-40.
WU Qiong, XIE XiangTing, WANG Lei, MOU Yong, LI JinWei. Development and Validation of Event-Specific PCR Method for the Quantification of Genetically Modified Soybean DBN8205[J]. Scientia Agricultura Sinica, 2026, 59(1): 29-40.
表1
DBN8205转化体和Lectin内标基因的引物/探针"
| 靶标 Target | 引物/探针 Primer/probe | 序列 Sequence (5′-3′) | 扩增子大小 Amplicon size (bp) | 来源 Reference | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| DBN8205 | DBN8205-QF | TTGAGGGGGTTACGCAATGTT | 120 | 自主研制 Self-developed | |||||||||||
| DBN8205-QR | GCGGGGGTCATAACGTGAC | ||||||||||||||
| DBN8205-QP | FAM-TCAGATTGTCGTTTCCCGCCTTCAG-BHQ | ||||||||||||||
| lectin | lectin-QF | GCCCTCTACTCCACCCCCA | 118 | [ | |||||||||||
| lectin-QR | GCCCATCTGCAAGCCTTTTT | ||||||||||||||
| lectin-QP | HEX-AGCTTCGCCGCTTCCTTCAACTTCAC-BHQ | ||||||||||||||
表3
DBN8205转化体特异性qPCR方法的定量限测试"
| 重复 Replicate | S5(100 copies, 0.25%) | S6(40 copies, 0.10%) | S7(20 copies, 0.05%) | S8 (10 copies, 0.025%) | ||||
|---|---|---|---|---|---|---|---|---|
| DBN8205拷贝数 DBN8205 copies (copies) | 百分比 Copy number ratio (%) | DBN8205拷贝数 DBN8205 copies (copies) | 百分比 Copy number ratio (%) | DBN8205拷贝数 DBN8205 copies (copies) | 百分比 Copy number ratio (%) | DBN8205拷贝数 DBN8205 copies (copies) | 百分比 Copy number ratio (%) | |
| 1 | 124 | 0.309 | 44 | 0.123 | 38 | 0.102 | 14 | 0.037 |
| 2 | 128 | 0.331 | 35 | 0.103 | 23 | 0.066 | 13 | 0.044 |
| 3 | 104 | 0.325 | 40 | 0.095 | 29 | 0.079 | 10 | 0.024 |
| 4 | 103 | 0.288 | 47 | 0.117 | 27 | 0.079 | 12 | 0.039 |
| 5 | 97 | 0.244 | 37 | 0.107 | 28 | 0.087 | 10 | 0.034 |
| 6 | 99 | 0.235 | 40 | 0.119 | 13 | 0.040 | 6 | 0.022 |
| 7 | 101 | 0.274 | 43 | 0.128 | 25 | 0.079 | 13 | 0.040 |
| 8 | 97 | 0.216 | 47 | 0.139 | 21 | 0.070 | 6 | 0.021 |
| 9 | 115 | 0.302 | 40 | 0.114 | 21 | 0.067 | 16 | 0.050 |
| 10 | 88 | 0.242 | 44 | 0.126 | 20 | 0.060 | 13 | 0.038 |
| 平均值 Average | 106 | 0.277 | 42 | 0.117 | 25 | 0.073 | 11 | 0.035 |
| 相对偏倚 Biasr (%) | 5.60 | 10.64 | 4.25 | 17.10 | 22.50 | 46.13 | 13.00 | 39.60 |
| 标准差 SD (%) | 12.74 | 0.04 | 4.00 | 0.01 | 3.30 | 0.01 | 3.30 | 0.010 |
| 相对标准差 RSD (%) | 12.06 | 14.66 | 9.60 | 11.08 | 29.22 | 27.80 | 29.22 | 27.80 |
表4
DBN8205转化体特异性定量PCR方法的准确性分析"
| 样品 Sample | 方法 Method | 预期含量 Expected content (%) | 重复 Replicate | 平均值 Average (%) | 相对偏倚 Biasr (%) | 标准差 SD (%) | 相对标准差 RSD (%) | P值 P value | ||
|---|---|---|---|---|---|---|---|---|---|---|
| Rep1 | Rep2 | Rep3 | ||||||||
| S1 | qPCR | 5.0 | 5.05 | 5.61 | 4.95 | 5.20 | 4.07 | 0.39 | 7.49 | 0.29 |
| ddPCR | 5.04 | 4.86 | 4.93 | 4.94 | -1.15 | 0.11 | 2.19 | |||
| S2 | qPCR | 3.0 | 3.20 | 2.93 | 2.91 | 3.01 | 0.44 | 0.17 | 5.68 | 0.95 |
| ddPCR | 3.04 | 2.93 | 3.05 | 3.01 | 0.24 | 0.07 | 2.35 | |||
| S3 | qPCR | 1.0 | 1.03 | 1.13 | 1.09 | 1.08 | 8.33 | 0.06 | 5.45 | 0.27 |
| ddPCR | 1.06 | 1.05 | 1.02 | 1.04 | 4.43 | 0.02 | 2.31 | |||
| S4 | qPCR | 0.5 | 0.50 | 0.55 | 0.55 | 0.53 | 6.67 | 0.03 | 5.54 | 0.19 |
| ddPCR | 0.52 | 0.49 | 0.50 | 0.50 | 0.85 | 0.01 | 2.75 | |||
| S6 | qPCR | 0.10 | 0.09 | 0.11 | 0.10 | 0.10 | 0.00 | 0.01 | 11.81 | 0.06 |
| ddPCR | 0.13 | 0.11 | 0.13 | 0.12 | 24.50 | 0.01 | 9.21 | |||
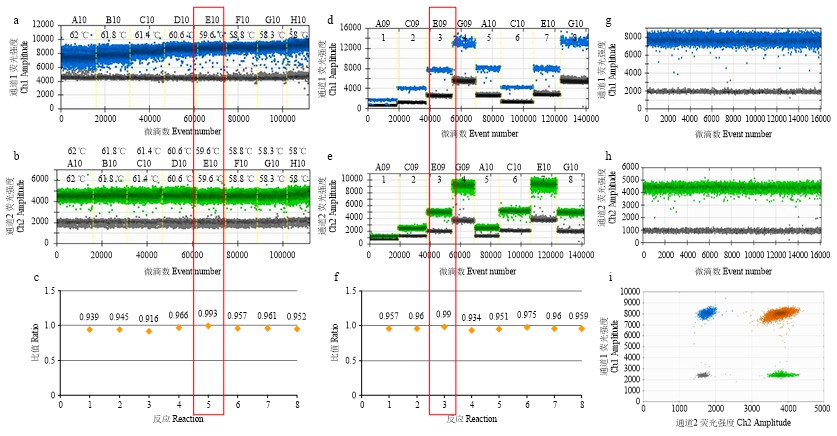
图2
DBN8205/Lectin二重微滴数字PCR引物/探针浓度、退火温度的优化 a:在58—62 ℃退火温度下DBN8205转化体一维(1-D)图;b:在58—62 ℃退火温度下退火温度优化中Lectin内标基因1-D图;c:不同退火温度下测量的DBN8205/Lectin拷贝数比值;d:引物/探针浓度优化中DBN8205转化体1-D图;e:引物/探针浓度优化中Lectin内标基因1-D图;f:不同引物/探针浓度下测量的DBN8205/Lectin拷贝数比值,1—4:DBN8205和Lectin的引物探针浓度相同,依次为100/50、200/100、400/200和800/400 nmol·L-1,5—8:DBN8205和Lectin的引物探针浓度不同,DBN8205引物/探针浓度依次为400/200、200/100、400/200和800/400 nmol·L-1,Lectin引物探针浓度依次为200/100、400/200、800/400和400/200 nmol·L-1;g:优化反应条件下DBN8205转化体1-D图;h:优化反应条件下Lectin内标基因的1-D图;i:优化的反应条件下DBN8205转化体和Lectin内标基因的2-D图"

表5
DBN8205/Lectin二重ddPCR测量梯度稀释DBN8205 DNA溶液的拷贝数浓度和比值"
| 靶标 Target | 预期模板量 Expected amount (copy/reaction) | 测量值 Measured value (copy/reaction ) | ||||||
|---|---|---|---|---|---|---|---|---|
| Rep1 | Rep2 | Rep3 | 均值 Mean | 标准差 SD | 相对标准差 RSD (%) | 相对偏倚 Biasr (%) | ||
| DBN8205 | 8.0×104 | 78000 | 77000 | 83000 | 79333 | 2624.67 | 3.31 | -0.83 |
| 1.0×104 | 10540 | 10680 | 10240 | 10487 | 183.55 | 1.75 | 4.87 | |
| 1.0×103 | 1384 | 1330 | 1430 | 1381 | 40.87 | 2.96 | 38.13 | |
| 500 | 506 | 502 | 504 | 504 | 1.63 | 0.32 | 0.80 | |
| 50 | 60 | 66 | 60 | 62 | 2.83 | 4.56 | 24.00 | |
| 40 | 46 | 40 | 52 | 46 | 4.90 | 10.65 | 15.00 | |
| Lectin | 8.0×104 | 79600 | 80000 | 86400 | 82000 | 3115.55 | 3.80 | 2.50 |
| 1.0×104 | 10800 | 10760 | 10700 | 10753 | 41.10 | 0.38 | 7.53 | |
| 1.0×103 | 1440 | 1428 | 1462 | 1443 | 14.08 | 0.98 | 44.33 | |
| 500 | 488 | 470 | 498 | 485 | 11.59 | 2.39 | -2.93 | |
| 50 | 56 | 56 | 58 | 57 | 0.94 | 1.66 | 13.33 | |
| 40 | 40 | 38 | 42 | 40 | 1.63 | 4.08 | 0.00 | |
| DBN8205/ Lectin | 8.0×104 | 0.98 | 0.96 | 0.96 | 0.97 | 0.01 | 0.90 | -3.23 |
| 1.0×104 | 0.98 | 0.99 | 0.96 | 0.98 | 0.01 | 1.49 | -2.48 | |
| 1.0×103 | 0.96 | 0.93 | 0.98 | 0.96 | 0.02 | 2.02 | -4.31 | |
| 500 | 1.04 | 1.07 | 1.01 | 1.04 | 0.02 | 2.21 | 3.90 | |
| 50 | 1.07 | 1.18 | 1.03 | 1.09 | 0.06 | 5.58 | 9.48 | |
| 40 | 1.15 | 1.05 | 1.24 | 1.15 | 0.08 | 6.60 | 14.69 | |
表6
8家实验室联合验证结果及统计分析"
| 样品 Samples | 预期值 Expected value (%) | ||||
|---|---|---|---|---|---|
| 5.0 | 3.0 | 1.0 | 0.5 | 0.1 | |
| 返回数据实验室数Labs reporting results | 8 | 8 | 8 | 8 | 8 |
| 每个实验室平行样品个数Replicate | 3 | 3 | 3 | 3 | 3 |
| 离群值数Replicate | 0 | 1 | 0 | 0 | 0 |
| 平均值Average (%) | 5.583 | 3.292 | 1.102 | 0.528 | 0.107 |
| 重复性标准差sr (%) | 0.201 | 0.117 | 0.109 | 0.036 | 0.009 |
| 重复性相对标准差RSDr (%) | 3.599 | 3.547 | 9.852 | 6.751 | 8.241 |
| 再现性标准差sR (%) | 0.337 | 0.177 | 0.124 | 0.046 | 0.008 |
| 再现性相对标准差RSDR (%) | 6.028 | 5.391 | 11.205 | 8.741 | 7.920 |
| 偏倚Bias (%) | 0.583 | 0.292 | 0.102 | 0.028 | 0.007 |
| 相对偏倚Biasr (%) | 11.660 | 9.742 | 10.224 | 5.695 | 7.297 |
| [1] |
AGBIOINVESTOR GM MONITOR. Global GM Crop Area Review. (2025-04-01) [2025-06-05]. https://gm.agbioinvestor.com.
|
| [2] |
doi: 10.3945/an.115.008870 pmid: 26567205 |
| [3] |
|
| [4] |
|
| [5] |
doi: 10.1021/acs.analchem.2c03680 pmid: 36205585 |
| [6] |
doi: 10.1007/s10068-023-01392-0 pmid: 38371688 |
| [7] |
doi: 10.1007/s00216-003-1767-7 |
| [8] |
pmid: 15123385 |
| [9] |
pmid: 23451388 |
| [10] |
doi: 10.1007/s00216-009-3150-9 pmid: 19789856 |
| [11] |
doi: 10.1007/s12010-017-2634-x |
| [12] |
doi: 10.1371/journal.pone.0002876 |
| [13] |
doi: 10.1093/clinchem/hvaa125 pmid: 32746458 |
| [14] |
doi: 10.1373/clinchem.2013.206375 pmid: 23570709 |
| [15] |
doi: 10.5740/jaoacint.16-0284 pmid: 28118137 |
| [16] |
doi: 10.1186/s13007-014-0042-6 pmid: 25628753 |
| [17] |
doi: 10.3390/foods13050747 |
| [18] |
doi: 10.1007/s00216-018-1010-1 pmid: 29574561 |
| [19] |
doi: 10.1016/j.jfca.2021.104236 |
| [20] |
pmid: 27990349 |
| [21] |
doi: 10.1007/s00216-019-01692-7 pmid: 30810790 |
| [22] |
doi: S0308-8146(19)30827-1 pmid: 31126507 |
| [23] |
INTERNATIONAL ORGANIZATION FOR STANDARDIZATION. Biotechnology requirements for evaluating the performance of quantification methods for nucleic acid target sequence-qPCR and dPCR. ISO 20395:2019, Geneva: International Organization for Standardization, 2019. https://www.iso.org/standard/67899.html.
|
| [24] |
doi: 10.1007/s00216-020-02834-y pmid: 32740822 |
| [25] |
全国农业转基因生物安全管理标准化技术委员会. 转基因植物及其产品成分检测大豆内标准基因定性PCR方法:农业部2031号公告-8-2013. 北京: 中国农业出版社, 2013: 1-6.
|
|
National Technical Committee for Standardization of Biosafety Management of Agricultural Genetically Modified Organisms. Detection of genetically modified plants and derived products-target-taxon- specific qualitative PCR method for soybean:Announcement by the Ministry of Agriculture No.2031-8-2013. Beijing: China Agriculture Press, 2013: 1-6. (in Chinese)
|
|
| [26] |
doi: 10.1007/BF02672069 |
| [27] |
|
| [28] |
Accuracy (trueness and precision) of measurement methods and results- Part 2: Basic method for the determination of repeatability and reproducibility of a standard measurement method: ISO 5725-2:2025. https://www.iso.org/standard/90054.html.
|
| [29] |
|
| [1] | 张琪, 陈二虎, 孙德宏, 唐培安. 锈赤扁谷盗谷胱甘肽S-转移酶基因CfGSTe1和CfGSTd1与甲酸乙酯耐受性关系[J]. 中国农业科学, 2026, 59(5): 1008-1019. |
| [2] | 江峰, 吴春燕, 王亦好, 杨泽众, 龚成, 罗晨. 烟粉虱MED脂肪酸延长酶基因家族鉴定和表达分析[J]. 中国农业科学, 2026, 59(4): 793-806. |
| [3] | 洪润婧, 周红, 蔺辉星, 范红结. 胞内劳森菌夹心ELISA检测方法的建立与应用[J]. 中国农业科学, 2025, 58(5): 1032-1042. |
| [4] | 王程泽, 张燕, 付伟, 贾京哲, 董金皋, 申珅, 郝志敏. 玉米ACO基因家族生物信息学及表达模式分析[J]. 中国农业科学, 2024, 57(7): 1308-1318. |
| [5] | 赵婕, 赵龙缘, 潘凝辉, 管丽蓉, 杜云龙, 李成云, 王云月, 谢勇. 水解酶基因BGIOSGA023826在稻瘟菌侵染过程中的抗病表型效应[J]. 中国农业科学, 2024, 57(23): 4607-4618. |
| [6] | 张英, 原青云, 任芳, 胡国君, 范旭东, 董雅凤. 葡萄浆果内坏死病毒RT-qPCR检测技术建立及其在葡萄砧木中的时空分布规律[J]. 中国农业科学, 2024, 57(14): 2771-2780. |
| [7] | 张小琴, 尹昌, 李政, 唐旭, 李艳, 吴春艳. 长期施肥对水稻土典型氨氧化菌和全程氨氧化菌种群活性和丰度的影响[J]. 中国农业科学, 2024, 57(14): 2803-2814. |
| [8] | 李俊, 单露英, 肖芳, 李允静, 高鸿飞, 翟杉杉, 吴刚, 张秀杰, 武玉花. 转基因玉米MON87427梯度含量基体标准物质的研制[J]. 中国农业科学, 2023, 56(8): 1444-1455. |
| [9] | 王晓阳, 彭振, 邢爱双, 赵盈睿, 马欣丽, 刘方, 杜雄明, 何守朴. 亚洲棉短纤维发育相关长链非编码RNA的鉴定及表达[J]. 中国农业科学, 2023, 56(23): 4565-4584. |
| [10] | 李俊, 赵新, 陈红, 李飞武, 梁晋刚, 李允静, 王颢潜, 高鸿飞, 张华, 陈子言, 吴刚, 沈平, 徐利群, 武玉花. 转基因定量检测结果测量不确定度的自上而下评定方法研究及应用[J]. 中国农业科学, 2023, 56(22): 4371-4385. |
| [11] | 李美璇, 张向昆, 王莉, 乔月莲, 师校欣, 杜国强. 沙地葡萄茎痘相关病毒在‘阳光玫瑰’葡萄树不同物候期和不同部位的变化规律[J]. 中国农业科学, 2023, 56(21): 4234-4244. |
| [12] | 苏佳, 赵炜, 刘丹, 王嘉, 白洪旭, 吴华伟, 薛青红, 陈晓春. 外源性马立克氏病病毒荧光定量PCR检测方法的建立[J]. 中国农业科学, 2023, 56(20): 4125-4136. |
| [13] | 曹鹏, 许建建, 李楚欣, 王新亮, 王春庆, 宋晨虎, 宋震. 柑橘黄化花叶病毒的实时定量PCR检测及其在寄主植株中的时空分布规律[J]. 中国农业科学, 2023, 56(18): 3574-3584. |
| [14] | 李允静, 肖芳, 武玉花, 李俊, 高鸿飞, 翟杉杉, 梁晋刚, 吴刚. 抗逆大豆IND-ØØ41Ø-5转化体特异性定量PCR检测方法的建立及其标准化[J]. 中国农业科学, 2023, 56(13): 2443-2460. |
| [15] | 翟晓虎, 李翎旭, 陈小竹, 蒋怀德, 贺卫华, 姚大伟. 肉中猪源性成分Real-time PCR定量检测技术[J]. 中国农业科学, 2023, 56(1): 156-164. |
|
||