













中国农业科学 ›› 2021, Vol. 54 ›› Issue (11): 2273-2286.doi: 10.3864/j.issn.0578-1752.2021.11.003
张林林( ),智慧,汤沙,张仁梁,张伟,贾冠清(
),智慧,汤沙,张仁梁,张伟,贾冠清( ),刁现民(
),刁现民( )
)
收稿日期:2020-11-09
接受日期:2020-12-07
出版日期:2021-06-01
发布日期:2021-06-09
联系方式:
张林林,E-mail:657044121@qq.com。
基金资助:
ZHANG LinLin( ),ZHI Hui,TANG Sha,ZHANG RenLiang,ZHANG Wei,JIA GuanQing(
),ZHI Hui,TANG Sha,ZHANG RenLiang,ZHANG Wei,JIA GuanQing( ),DIAO XianMin(
),DIAO XianMin( )
)
Received:2020-11-09
Accepted:2020-12-07
Published:2021-06-01
Online:2021-06-09
摘要:
【目的】谷子抽穗时间的适应性表现是广适性新品种选育的基础,分析抽穗时间关键基因的遗传变异和单倍型效应,为品种适应性改良提供基础信息。【方法】通过全基因组关联分析(genome-wide association study,GWAS),定位谷子抽穗时间关键基因SiTOC1,利用多组学数据库(multi-omics database for Setaria italica,MDSi)提供的SiTOC1数字表达量,分析SiTOC1的组织时空表达特性,并利用原生质体对SiTOC1蛋白进行亚细胞定位。采用qRT-PCR在短日(10 h光照/14 h黑暗)条件下进行SiTOC1 24 h节律表达模式分析。利用有代表性的99份谷子品种,分析SiTOC1编码区和启动子区的遗传多态性、单倍型以及转录水平,并对单倍型与抽穗时间的关系进行鉴定。【结果】在第1染色体物理位置31 456 761 bp处鉴定到了一个显著的关联信号,与抽穗时间紧密相关,该位点附近存在一个拟南芥抽穗期TOC1的同源基因SiTOC1。SiTOC1在光周期响应组织(根、茎、叶等)中高表达,亚细胞定位于细胞核,在傍晚表达量上调,呈现出24 h节律性表达模式。SiTOC1在不同谷子品种中存在丰富的多态性,但REC和CCT结构域高度保守。SiTOC1编码区2种主要单倍型H-2和H-6分别与启动子单倍型Hp-591C和Hp-591A共分离,其中,启动子单倍型Hp-591C较Hp-591A的相对表达量显著上调了约2.5倍(P=0.014),并且该单倍型在三亚市、长治市和乌鲁木齐市3个环境下的抽穗时间分别平均延迟9、11和12 d。【结论】SiTOC1启动子区第591 bp处的SNP是引起抽穗时间差异的主效位点,单倍型Hp-591A较Hp-591C早熟,可作为主效单倍型用于分子育种选择。
张林林,智慧,汤沙,张仁梁,张伟,贾冠清,刁现民. 谷子抽穗时间基因SiTOC1的表达与单倍型变异分析[J]. 中国农业科学, 2021, 54(11): 2273-2286.
ZHANG LinLin,ZHI Hui,TANG Sha,ZHANG RenLiang,ZHANG Wei,JIA GuanQing,DIAO XianMin. Characterizations of Transcriptional and Haplotypic Variations of SiTOC1 in Foxtail Millet[J]. Scientia Agricultura Sinica, 2021, 54(11): 2273-2286.
表1
SiTOC1亚细胞定位载体构建PCR扩增及测序引物"
| 载体 Vector | 引物名称 Primer name | 引物序列 Primer sequence (5′-3′) | 产物长度 Product length (bp) |
|---|---|---|---|
| GFP-N 端载体 GFP-N terminal vector | GFPG5-F1 | GCTGTACAAGACTAGTATGGTGGGCGGCGGCG | 1517 |
| GFPG5-R1 | GGGGAAATTCGAGCTCCTACTCTGGAGAAGAAATAATC | ||
| PUCGFPN-TestF | TACAACTACAACAGCCACAA | 2163 | |
| PUCGFPN-TestR | CCTCTTCGCTATTACGC | ||
| GFPG5-F1.1 | CCATCATCATGATGTCC | ||
| GFPG5-R1.1 | TTCAGAGCGACTGCATG | ||
| GFP-C端载体 GFP-C terminal vector | G5GFP-F1 | TAGTGGATCCATCGATATGGTGGGCGGCGGCG | 1514 |
| G5GFP-R1 | TCCCGGGAGCGGTACCCTCTGGAGAAGAAATAATCTC | ||
| PUCGFPC-TestF | ACCTCCTCGGATTCCAT | 2354 | |
| PUCGFPC-TestR | TGCCGTTCTTCTGCTTG | ||
| G5GFP-F1.1 | CAAGATGCTCAAGTACA | ||
| G5GFP-R1.1 | CGATTTACATACCTCACT | ||
| RFP-N端载体 RFP-N terminal vector | MYBRFP-F | GCTGTACAAGACTAGTATGGACATGGCGCACGAGAG | 1022 |
| MYBRFP-R | GGGGAAATTCGAGCTCTCACACGGCGGCCTGGGT | ||
| PUCRFPN-TestF | CTCAAGCTCAAGGACGG | 1584 | |
| PUCRFPN-TestR | CCTCTTCGCTATTACGC |
表2
SiTOC1 PCR扩增和测序引物"
| 产物Product | 引物名称 Primer name | 引物序列 Primer sequence (5′-3′) | 产物长度 Product length (bp) |
|---|---|---|---|
| 编码区 Coding region | G5-F1 | GATCCAGCGACAGTCCA | 1450 |
| G5-R1 | AGACAGGTCGGACTGAAATA | ||
| G5-F2 | GATGAGTTGCCACAAAGATG | 1272 | |
| G5-R2 | TGTATTGCCTTTCCCAGTAG | ||
| 启动子区 Promoter region | G5P-F1 | GGCCAAACGAAACCATG | 2069 |
| G5P-R1 | CCTAATCCGGGAAACCAG | ||
| G5P-F1.1 | ATTGCGTGCACGAATCT | ||
| G5P-R1.1 | CAGCAGCAGCCTGCCTCG | ||
| SiTOC1互补DNA SiTOC1 cDNA | G5Q-F1 | GCCGATCAAGCATCATATGTTAAGT | 250 |
| G5Q-R1 | TTTGGCCTTCATTGCTTCGC | ||
| Cullin互补DNA Cullin cDNA | Cullin-F | TATGGGTCATCAACAGCTTGTC | 112 |
| Cullin-R | GTAGTCCCTCGTGATGAGATCC |
表3
候选基因注释信息"
| 位点名称 LocusName | 拟南芥对应基因 Arabi-symbol | 拟南芥基因功能注释 Arabi-defline |
|---|---|---|
| Seita.1G235800 | N/A | 无功能注释N/A |
| Seita.1G235900 | MKM21.12 | 线粒体核糖体蛋白L27 Mitochondrial ribosomal protein L27 |
| Seita.1G236000 | N/A | 谷氧还蛋白家族 Glutaredoxin family protein |
| Seita.1G236100 | TOC1 | 含CCT基序的应答调节蛋白 CCT motif-containing response regulator protein |
| Seita.1G236200 | PAP10 | 紫色酸性磷酸酶10 Purple acid phosphatase 10 |
| Seita.1G236300 | NOP10 | 核仁RNA结合Nop10p家族蛋白 Nucleolar RNA-binding Nop10p family protein |
| Seita.1G236400 | AtMYB79 | 含MYB结构域蛋白79 MYB domain protein 79 |
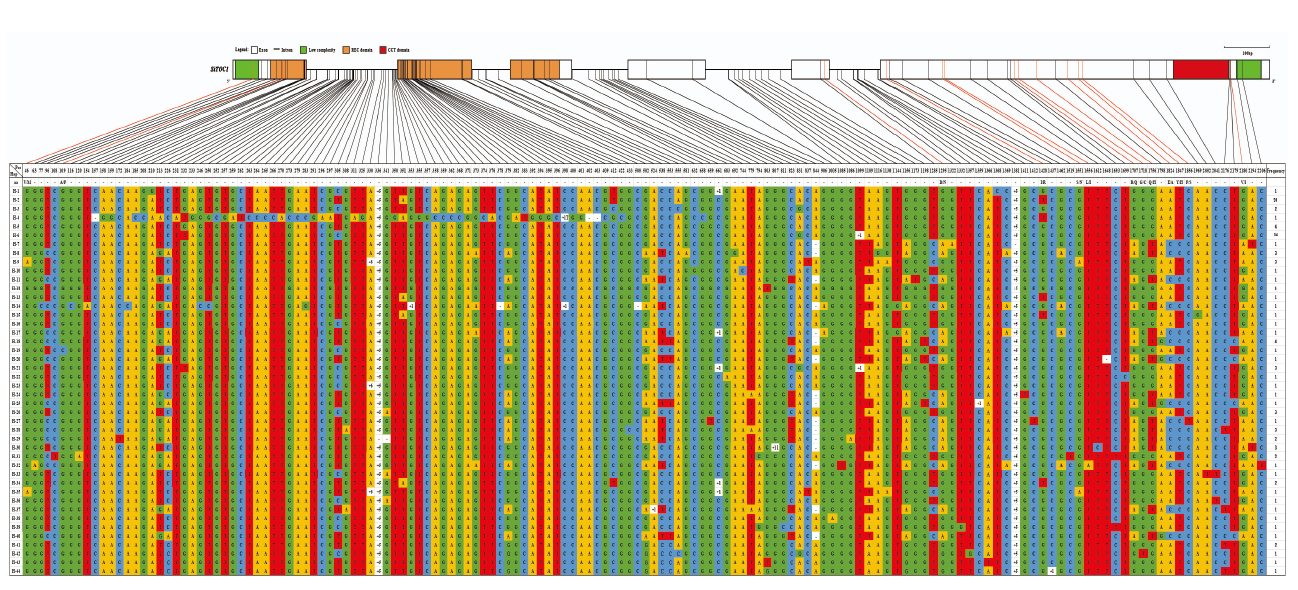
图5
SiTOC1编码区结构变异 图上方为SiTOC1编码区基因结构,其中,框格代表外显子,框格之间的连接线代表内含子,下方对应不同单倍型组合信息表,连接上方SiTOC1编码区基因结构图的实心线指示变异位点,其中,红色连接线指示非同义突变,表格中不同颜色单元格代表不同碱基,其中,黄色、红色、蓝色和绿色单元格分别代表A碱基、T碱基、C碱基、G碱基,-:缺失,+:插入,符号后面的数字代表缺失或插入碱基数,Indel位点从左往右依次为:330(+6)/+gtgctg、336(+5)/+atgct、353(+1)/+t、398(+1)/+t、524(+1)/+t、661(+1)/+g、807(+11)/+gtgggttgctt、1099(+1)/+c、1359(+1)/+t、1381(+5)/+taatc、1437(+1)/+t"

| [1] | TILMAN D, BALZER C, HILL J, BEFORT B L. Global food demand and the sustainable intensification of agriculture. Proceedings of the National Academy of Sciences of the United States of America, 2011,108(50):20260-20264. |
| [2] | BARTON L, NEWSOME S D, CHEN F H, WANG H, GUILDERSON T P, BETTINGER R L. Agricultural origins and the isotopic identity of domestication in northern China. Proceedings of the National Academy of Sciences of the United States of America, 2009,106(14):5523-5528. |
| [3] |
JIA G Q, HUANG X H, ZHI H, ZHAO Y, ZHAO Q, LI W J, CHAI Y, YANG L, LIU K Y, LU H Y, ZHU C R, LU Y Q, ZHOU C C, FAN D L, WENG Q J, GUO Y L, HUANG T, ZHANG L, LU T T, FENG Q, HAO H F, LIU H K, LU P, ZHANG N, LI Y H, GUO E, WANG S J, WANG S Y, LIU J R, ZHANG W F, CHEN G Q, ZHANG B G, LI W, WANG Y F, LI H Q, ZHAO B H, LI J Y, DIAO X M, HAN B. A haplotype map of genomic variations and genome-wide association studies of agronomic traits in foxtail millet (Setaria italica). Nature Genetics, 2013,45(8):957-961.
doi: 10.1038/ng.2673 |
| [4] |
BENNETZEN J L, SCHMUTZ J, WANG H, PERCIFIELD R, HAWKINS J, PONTAROLI A C, ESTEP M, FENG L, VAUGHN J N, GRIMWOOD J, JENKINS J, BARRY K, LINDQUIST E, HELLSTEN U, DESHPANDE S, WANG X W, WU X M, MITROS T, TRIPLETT J, YANG X H, YE C Y, MAURO-HERRERA M, WANG L, LI P H, SHARMA M, SHARMA R, RONALD P C, PANAUD O, KELLOGG E A, BRUTNELL T P, DOUST A N, TUSKAN G A, ROKHSAR D, DEVOS K M. Reference genome sequence of the model plant Setaria. Nature Biotechnology, 2012,30(6):555-561.
doi: 10.1038/nbt.2196 |
| [5] |
ZHANG G Y, LIU X, QUAN Z W, CHENG S F, XU X, PAN S K, XIE M, ZENG P, YUE Z, WANG W L, TAO Y, BIAN C, HAN C L, XIA Q J, PENG X H, CAO R, YANG X H, ZHAN D L, HU J C, ZHANG Y X, LI H N, LI H, LI N, WANG J Y, WANG C C, WANG R Y, GUO T, CAI Y J, LIU C Z, XIANG H T, SHI Q X, HUANG P, CHEN Q C, LI Y R, WANG J, ZHAO Z H, WANG J. Genome sequence of foxtail millet (Setaria italica) provides insights into grass evolution and biofuel potential. Nature Biotechnology, 2012,30(6):549-554.
doi: 10.1038/nbt.2195 |
| [6] |
YANG Z R, ZHANG H S, LI X K, SHEN H M, GAO J H, HOU S Y, ZHANG B, MAYES S, BENNETT M, MA J X, WU C Y, SUI Y, HAN Y H, WANG X C. A mini foxtail millet with an Arabidopsis-like life cycle as a C4 model system. Nature Plants, 2020,6(9):1167-1178.
doi: 10.1038/s41477-020-0747-7 |
| [7] |
COLASANTI J, CONEVA V. Mechanisms of floral induction in grasses: Something borrowed, something new. Plant Physiology, 2009,149(1):56-62.
doi: 10.1104/pp.108.130500 |
| [8] |
ANDRÉS F, COUPLAND G. The genetic basis of flowering responses to seasonal cues. Nature Review Genetics, 2012,13(9):627-639.
doi: 10.1038/nrg3291 |
| [9] |
RIESEBERG L H, WILLIS J H. Plant speciation. Science, 2007,317(5840):910-914.
doi: 10.1126/science.1137729 |
| [10] |
STRAYER C, OYAMA T, SCHULTZ T F, RAMAN R, SOMERS D E, MÁS P, PANDA S, KREPS J A, KAY S A. Cloning of the Arabidopsis clock gene TOC1, an autoregulatory response regulator homolog. Science, 2000,289(5480):768-771.
doi: 10.1126/science.289.5480.768 |
| [11] |
COCKRAM J, THIEL T, STEUERNAGEL B, STEIN N, TAUDIEN S, BAILEY P C, O'SULLIVAN D M. Genome dynamics explain the evolution of flowering time CCT domain gene families in the Poaceae. PLoS ONE, 2012,7(9):e45307.
doi: 10.1371/journal.pone.0045307 |
| [12] |
WENKEL S, TURCK F, SINGER K, GISSOT L, LE GOURRIEREC J L, SAMACH A, COUPLAND G. CONSTANS and the CCAAT box binding complex share a functionally important domain and interact to regulate flowering of Arabidopsis. The Plant Cell, 2006,18(11):2971-2984.
doi: 10.1105/tpc.106.043299 |
| [13] | GENDRON J M, PRUNEDA-PAZ J L, DOHERTY C J, GROSS A M, KANG S E, KAY S A. Arabidopsis circadian clock protein, TOC1, is a DNA-binding transcription factor. Proceedings of the National Academy of Sciences of the United States of America, 2012,109(8):3167-3172. |
| [14] |
MAKINO S, MATSUSHIKA A, KOJIMA M, YAMASHINO T, MIZUNO T. The APRR1/TOC1 quintet implicated in circadian rhythms of Arabidopsis thaliana: I. Characterization with APRR1- overexpressing plants. Plant Cell Physiology, 2002,43(1):58-69.
doi: 10.1093/pcp/pcf005 |
| [15] |
SHIM J S, KUBOTA A, IMAIZUMI T. Circadian clock and photoperiodic flowering in Arabidopsis: CONSTANS is a hub for signal integration. Plant Physiology, 2017,173(1):5-15.
doi: 10.1104/pp.16.01327 |
| [16] |
KOO B H, YOO S C, PARK J W, KWON C T, LEE B D, AN G, ZHANG Z, LI J, LI Z, PAEK N C. Natural variation in OsPRR37 regulates heading date and contributes to rice cultivation at a wide range of latitudes. Molecular Plant, 2013,6(6):1877-1888.
doi: 10.1093/mp/sst088 |
| [17] |
SALOMÉP A, MCCLUNG C R. The Arabidopsis thaliana clock. Journal of Biological Rhythms, 2004,19(5):425-435.
doi: 10.1177/0748730404268112 |
| [18] |
MÁS P. Circadian clock signaling in Arabidopsis thaliana: From gene expression to physiology and development. The International Journal of Developmental Biology, 2005,49(5/6):491-500.
doi: 10.1387/ijdb.041968pm |
| [19] |
GARDNER M J, HUBBARD K E, HOTTA C T, DODD A N, WEBB A A. How plants tell the time. Biochemical Journal, 2006,397(1):15-24.
doi: 10.1042/BJ20060484 |
| [20] |
ALABADÍ D, OYAMA T, YANOVSKY M J, HARMON F G, MÁS P, KAY S A. Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock. Science, 2001,293(5531):880-883.
doi: 10.1126/science.1061320 |
| [21] |
PRUNEDA-PAZ J L, BRETON G, PARA A, KAY S A. A functional genomics approach reveals CHE as a component of the Arabidopsis circadian clock. Science, 2009,323(5920):1481-1485.
doi: 10.1126/science.1167206 |
| [22] | 陆平. 谷子种质资源描述规范和数据标准2-9. 北京: 中国农业出版社, 2006. |
| LU P. Description Specification and Data Standard of Foxtail Millet Germplasm Resources 2-9. Beijing: China Agriculture Press, 2006. (in Chinese) | |
| [23] | TURNER S D. qqman: An R package for visualizing GWAS results using Q-Q and manhattan plots. biorXiv, 2014, https://doi.org/10.1101/005165. |
| [24] |
YANG A, DAI X Y, ZHANG W H. A R2R3-type MYB gene, OsMYB2, is involved in salt, cold, and dehydration tolerance in rice. Journal of Experimental Botany, 2012,63(7):2541-2556.
doi: 10.1093/jxb/err431 |
| [25] |
LIVAK K J, SCHMITTGEN T D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods, 2001,25(4):402-408.
doi: 10.1006/meth.2001.1262 |
| [26] | DOYLE J. DNA protocols for plants-CTAB total DNA isolation. Molecular Techniques in Taxonomy, 1991: 283-293. |
| [27] |
LIBRADO P, ROZAS J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics, 2009,25(11):1451-1452.
doi: 10.1093/bioinformatics/btp187 |
| [28] | 刁现民, 程汝宏. 十五年区试数据分析展示谷子糜子育种现状. 中国农业科学, 2017,50(23):4469-4474. |
| DIAO X M, CHENG R H. Fifteen-year regional trial data analysis shows the current situation of millet and millet breeding. Scientia Agricultura Sinica, 2017,50(23):4469-4474. (in Chinese) | |
| [29] |
YANO M, KATAYOSE Y, ASHIKARI M, YAMANOUCHI U, MONNA L, FUSE T, BABA T, YAMAMOTO K, UMEHARA Y, NAGAMURA Y, SASAKI T. Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. The Plant Cell, 2000,12(12):2473-2484.
doi: 10.1105/tpc.12.12.2473 |
| [30] |
HAYAMA R, YOKOI S, TAMAKI S, YANO M, SHIMAMOTO K. Adaptation of photoperiodic control pathways produces short-day flowering in rice. Nature, 2003,422(6933):719-722.
doi: 10.1038/nature01549 |
| [31] |
XUE W Y, XING Y Z, WENG X Y, ZHAO Y, TANG W J, WANG L, ZHOU H J, YU S B, XU C G, LI X H, ZHANG Q F. Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nature Genetics, 2008,40(6):761-767.
doi: 10.1038/ng.143 |
| [32] |
NEMOTO Y, NONOUE Y, YANO M, IZAWA T. Hd1, a CONSTANS ortholog in rice, functions as an Ehd1 repressor through interaction with monocot-specific CCT-domain protein Ghd7. The Plant Journal, 2016,86(3):221-233.
doi: 10.1111/tpj.2016.86.issue-3 |
| [33] |
DU A, TIAN W, WEI M H, YAN W, HE H, ZHOU D, HUANG X, LI S G, OUYANG X H. The DTH8-Hd1 module mediates day-length- dependent regulation of rice flowering. Molecular Plant, 2017,10(7):948-961.
doi: 10.1016/j.molp.2017.05.006 |
| [34] |
FUJINO K, YAMANOUCHI U, NONOUE Y, OBARA M, YANO M. Switching genetic effects of the flowering time gene Hd1 in LD conditions by Ghd7 and OsPRR37 in rice. Breeding Science, 2019,69(1):127-132.
doi: 10.1270/jsbbs.18060 |
| [35] |
ZHANG Z Y, ZHANG B, QI F X, WU H, LI Z X, XING Y Z. Hd1 function conversion in regulating heading is dependent on gene combinations of Ghd7, Ghd8, and Ghd7.1 under long-day conditions in rice. Molecular Breeding, 2019,39(92):1-12.
doi: 10.1007/s11032-018-0907-x |
| [1] | 叶美金, 吴雷, Lohani Md Nahibuzzaman, 尹丽, 胡欣荣, 刘亚西, 蒋云峰, 陈国跃, 蒲至恩, 李阳, 李婷, 邹亚亚, 吴佳怡, 马建. 基于GWAS的中国地方小麦成熟胚大小位点的鉴定及其遗传效应解析[J]. 中国农业科学, 2026, 59(6): 1157-1171. |
| [2] | 杨丽娟, 陈丝雨, 赵薇, 朱玲, 郭磊, 马丽娜, 马瑞敏, 张娟. 全基因组重测序揭示静原鸡羽色的遗传机制[J]. 中国农业科学, 2026, 59(6): 1348-1360. |
| [3] | 王勇胜, 牛丽, 王长杰, 马立花, 廉潇潇, 孟亚雄, 马小乐, 姚立蓉, 张宏, 杨轲, 李葆春, 王化俊, 司二静, 汪军成. 冬小麦千粒重的全基因组关联分析及候选基因预测[J]. 中国农业科学, 2026, 59(3): 499-514. |
| [4] | 张义茹, 韩雪, 姚鑫杰, 冯军, 魏爱丽, 李文超, 张彬, 韩渊怀, 李红英. 基于多组学解析谷子后熟米色变化的分子机制[J]. 中国农业科学, 2025, 58(9): 1702-1718. |
| [5] | 李云丽, 刁邓超, 刘雅睿, 孙玉晨, 孟祥宇, 邬陈芳, 汪妤, 吴建辉, 李春莲, 曾庆东, 韩德俊, 郑炜君. 小麦苗期耐热性全基因组关联分析[J]. 中国农业科学, 2025, 58(9): 1663-1683. |
| [6] | 周广飞, 马亮, 马璐, 张舒钰, 章慧敏, 宋旭东, 张振良, 陆虎华, 郝德荣, 冒宇翔, 薛林, 陈国清. 玉米苞叶性状全基因组关联分析[J]. 中国农业科学, 2025, 58(3): 431-442. |
| [7] | 武书羽, 衡燕芳, 于太飞, 王世佳, 于思佳, 李园, 胡正, 张辉, 孙现军, 黎亮, 姜奇彦. 玉米自然群体苗期耐盐性鉴定及耐盐相关基因分析[J]. 中国农业科学, 2025, 58(20): 4085-4099. |
| [8] | 李明, 程宇坤, 白斌, 雷斌, 耿洪伟. 冬小麦穗部性状GWAS分析及优异单倍型筛选[J]. 中国农业科学, 2025, 58(18): 3583-3597. |
| [9] | 向爱慧, 白荣基, 郝宇琼, 赵佳佳, 武棒棒, 李晓华, 郑兴卫, 关攀锋, 郑军. 山西小麦矮秆基因的鉴定及株高遗传位点挖掘[J]. 中国农业科学, 2025, 58(17): 3372-3388. |
| [10] | 郑敏华, 陈洛, 邢甲乐, 谢月兰, 姜先芽, 聂帅, 蔡甫格, 巫浩翔, 陆展华, 孙伟, 霍兴, 白嵩, 赵均良, 杨武. 华南籼稻稻瘟病抗性QTL鉴定与候选基因挖掘[J]. 中国农业科学, 2025, 58(14): 2707-2719. |
| [11] | 李宁, 高丽锋, 黄鑫, 史华伟, 杨进文, 史雨刚, 陈明, 贾继增, 孙黛珍. 耐低氮小麦品种的筛选及耐低氮指数的全基因组关联分析[J]. 中国农业科学, 2025, 58(13): 2487-2503. |
| [12] | 史顺宇, 杨涛, 庞博, 李静, 林轶峰, 王正瑞, 傅林成, 扎尔加玛丽·阿不都别克, 高文伟, 吴鹏昊. 海岛棉叶绿素含量的全基因组关联分析及候选基因预测[J]. 中国农业科学, 2025, 58(10): 1878-1895. |
| [13] | 张颖, 石婷瑞, 曹瑞, 潘文秋, 宋卫宁, 王利, 聂小军. ICARDA引进-小麦苗期抗旱性的全基因组关联分析[J]. 中国农业科学, 2024, 57(9): 1658-1673. |
| [14] | 赵真坚, 王凯, 陈栋, 申琦, 余杨, 崔晟頔, 王俊戈, 陈子旸, 禹世欣, 陈佳苗, 王翔枫, 唐国庆. 基因组和DNA甲基化组联合分析筛选猪肉质性状关键基因[J]. 中国农业科学, 2024, 57(7): 1394-1406. |
| [15] | 周浩露, 申朝阳, 罗新宇, 黄英惠, 王可心, 王云浩, 高小丽. 氮肥对垄沟集雨种植谷子氮素利用效率及产量的影响[J]. 中国农业科学, 2024, 57(5): 885-899. |
|
||