













中国农业科学 ›› 2023, Vol. 56 ›› Issue (10): 1966-1981.doi: 10.3864/j.issn.0578-1752.2023.10.012
王霏1( ), 肖迎珂1, 宣旭娴1, 张晓雯1, 刘菲1, 查紫仙1, 代梦瞳1, 王西成2, 吴伟民2, 房经贵1, 王晨1(
), 肖迎珂1, 宣旭娴1, 张晓雯1, 刘菲1, 查紫仙1, 代梦瞳1, 王西成2, 吴伟民2, 房经贵1, 王晨1( )
)
收稿日期:2022-07-04
接受日期:2022-12-27
出版日期:2023-05-16
发布日期:2023-05-17
通信作者:
王晨,E-mail:wangchen@njau.edu.cn
联系方式:
王霏,E-mail:2020104031@stu.njau.edu.cn。
基金资助:
WANG Fei1( ), XIAO YingKe1, XUAN XuXian1, ZHANG XiaoWen1, LIU Fei1, ZHA ZiXian1, DAI MengTong1, WANG XiCheng2, WU WeiMin2, FANG JingGui1, WANG Chen1(
), XIAO YingKe1, XUAN XuXian1, ZHANG XiaoWen1, LIU Fei1, ZHA ZiXian1, DAI MengTong1, WANG XiCheng2, WU WeiMin2, FANG JingGui1, WANG Chen1( )
)
Received:2022-07-04
Accepted:2022-12-27
Published:2023-05-16
Online:2023-05-17
摘要:
【目的】鉴定葡萄miR164s(VvmiR164a/b/c/d)及其靶基因,明确VvmiR164s及其靶基因应答外源赤霉素在葡萄单性结实过程中的调控作用。【方法】以‘魏可’葡萄(Vitis vinifera L. Wink)为试材,利用miR-RACE、PCR、RLM-RACE与PPM-RACE、qRT-PCR及生物信息学等技术,分析VvmiR164s-VvNAC100模块应答外源赤霉素在葡萄单性结实过程中的时空表达特征及其潜在功能。【结果】赤霉素在花前处理‘魏可’葡萄,可诱导其单性结实,导致葡萄的无核。克隆鉴定了VvmiR164a/b/c/d的精确序列,预测到其4条靶基因VvNAC100-1、VvNAC100-2、VvNAC098、VvNAC021,结合匹配程度与前期数据,在此重点分析靶基因VvNAC100-2,并将其命名为VvNAC100。克隆鉴定靶基因VvNAC100,验证其裂解位点,并对其开展蛋白进化、染色体定位及结构分析。VvNAC100裂解位点位于miRNA 5′端的第9位与第11位,裂解频度为17/20与11/20,定位于葡萄的Chr14上,编码363个氨基酸,含有一个NAM结构域,且其定位于细胞核。VvNAC100蛋白与其他物种氨基酸序列保守性较高,功能相似,其中与辣椒、烟草等物种亲缘关系较近。VvMIR164a/b/c/d的启动子与其靶基因VvNAC100的启动子均包含多种激素作用元件,表明其可能通过响应不同的激素来参与葡萄生长发育的调控。RT-qPCR结果显示,随着葡萄子房的发育,VvmiR164b的表达水平呈下降趋势,其靶基因VvNAC100在子房发育前期呈上升的表达趋势,具有一定的负相关,而VvmiR164a/c/d与VvNAC100表达模式相似,呈现一定的正相关;GA处理后,VvmiR164a/c/d在葡萄子房单性结实过程中的表达极显著地上升,进而也显著抑制了VvNAC100在这一时期的表达,从而促进了VvmiR164a/c/d-VvNAC100表达水平的负相关;但VvmiR164b在GA处理后表现出下降的趋势,且其与VvNAC100的表达水平呈现一定的正相关,表明GA处理增强了VvmiR164a/c/d对VvNAC100的负调控,减弱了VvmiR164b对VvNAC100的负调控作用。【结论】VvNAC100是VvmiR164 家族VvmiR164a/b/c/d的真实靶基因;在葡萄单性结实过程中,赤霉素诱导了VvmiR164a/c/d对靶基因VvNAC100的负调控作用,抑制了VvmiR164b对靶基因VvNAC100的负调控作用,VvmiR164a/c/d是VvmiR164家族参与赤霉素诱导葡萄单性结实的主效因子。
王霏, 肖迎珂, 宣旭娴, 张晓雯, 刘菲, 查紫仙, 代梦瞳, 王西成, 吴伟民, 房经贵, 王晨. VvmiR164s-VvNAC100作用模块鉴定及其在葡萄子房发育过程中响应赤霉素的表达分析[J]. 中国农业科学, 2023, 56(10): 1966-1981.
WANG Fei, XIAO YingKe, XUAN XuXian, ZHANG XiaoWen, LIU Fei, ZHA ZiXian, DAI MengTong, WANG XiCheng, WU WeiMin, FANG JingGui, WANG Chen. Identification of the VvmiR164s-VvNAC100 Action Module and Analysis of Their Expressions Responsive to Exogenous GA During Grape Ovary Development[J]. Scientia Agricultura Sinica, 2023, 56(10): 1966-1981.
表1
VvmiR164s PCR 扩增引物序列"
| 基因名称 Gene name | 正向引物序列 Forward primer sequence | 反向引物序列 Reverse primer sequence | 用途 Application |
|---|---|---|---|
| VvmiR164aQT | GATGAATTGCCTACAGTTACCAG | CCAAGGCCTCCTGCAATTT | VvmiR164a前体序列的扩增 Amplification of VvmiR164a precursor |
| VvmiR164bQT | GATGGTCAGCTGCCTAATATTG | ACACGCATCTTGGATTTGCT | VvmiR164b前体序列的扩增 Amplification of VvmiR164b precursor |
| VvmiR164cQT | CTTCATTTGCCGCTTTCG | CTCCACCAAGAAAATGATAGGC | VvmiR164c前体序列的扩增 Amplification of VvmiR164c precursor |
| VvmiR164dQT | GCAATCACTTGGGAGTTGG | ATAGCCAATAGAGGGGAAAAGG | VvmiR164d前体序列的扩增 Amplification of VvmiR164d precursor |
| VvNAC100 | ATGGAAAAGGCTCCTGACCA | GGAATGTTTCCCTTCTAATCTGT | VvNAC100靶全长序列的扩增 Amplification of full-length sequences of VvNAC100 |
| VvNAC100p | ATGGAAAAGGCTCCTGACC | GTAATTCCAGAGGCAATCAATG | VvNAC100靶片段序列的扩增 Amplification of VvNAC100 target fragment sequences |
| VvmiR164a-qRT | TCAAAACCCGAAACAAGTCC | ACCCACCCTCAACCATACAA | VvmiR164a定量RT-PCR引物 Real-time PCR of VvmiR164a |
| VvmiR164b-qRT | CGAGTGTTGAGCAAGATGGA | CTCCGTGATAATTGGGGAGA | VvmiR164b定量RT-PCR引物 Real-time PCR of VvmiR164b |
| VvmiR164c-qRT | TTGGGGGTGTGAAGAAGAAG | GAGGAACCCATGTTGGAGAA | VvmiR164c 定量RT-PCR引物 Real-time PCR of VvmiR164c |
| VvmiR164d-qRT | TTGGGGGTGTGAAGAAGAAG | GAGGAACCCATGTTGGAGAA | VvmiR164d定量RT-PCR引物 Real-time PCR of VvmiR164d |
| VvNAC100-qRT | TTCTCATCCCTGACCAGTCC | TCTACAGGCCCAGCTGAAGT | VvNAC100定量RT-PCR引物 Real-time PCR of VvNAC100 |
| Vvactin | GCTCGCTGTTTTGCAGTTCTAC | AACATAGGTGAGGCCGCACTT | 葡萄actin内参基因 Actin internal reference primer |
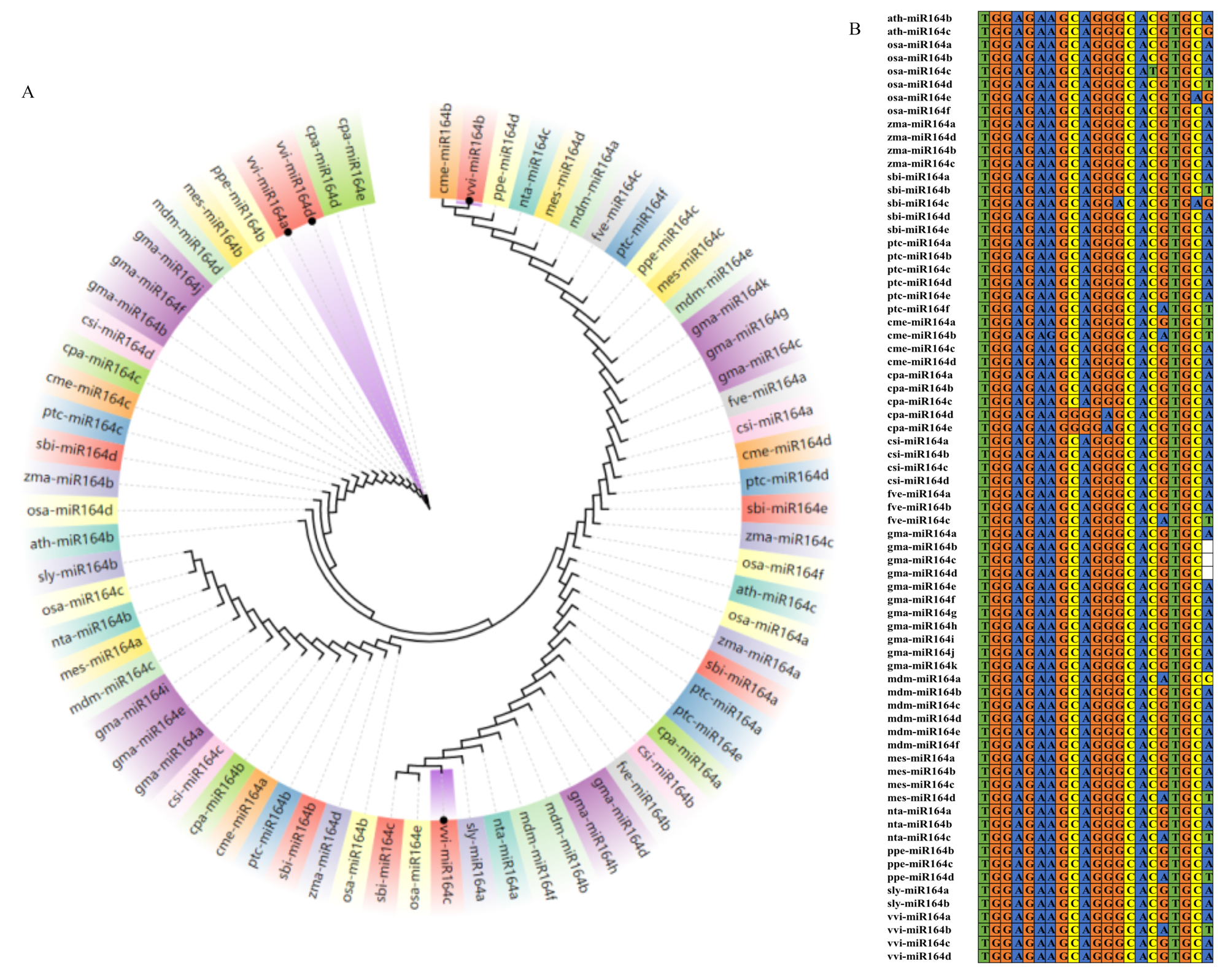
图3
VvmiR164s进化分析(A)和成熟体序列比对(B) ath:拟南芥Arabidopsis thaliana;cme:甜瓜Cucumis melo;cpa:番木瓜Carica papaya;csi:柑橘Citrus sinensis;fve:草莓Fragaria vesca;gma:大豆Glycine max;mdm:苹果Malus domestica;nta:烟草Nicotiana tabacum;osa:水稻Oryzasativa;ppe:桃Prunus persica;ptc:毛果杨 Populus trichocarpa;sbi:高粱Sorghum bicolor;sly:番茄Solanum lycopersicum;mes:木薯Manihot esculenta;zma:玉米Zea mays;vvi:葡萄Vvtis vinifera"

表2
葡萄VvmiR164s及靶基因相关信息"
| miRNAs | MiRNA 成熟体长度 Length of miRNA | 靶基因编号 Target_Acc. | 错配率Expectation | 靶区 Target region | 靶基因匹配 初始位置 Target_start | 靶基因匹配 初始位置 Target_end | 切割方式 Interaction mode inhibition |
|---|---|---|---|---|---|---|---|
| VvmiR164a/c/d (UGGAGAAGCAGGGCACGUGCA) | 21 | VIT_217s0000g06400 (VvNAC100-1) | 0.5 | gene=VIT_217s0000g06400 CDS=732-1814 | 1375 | 1395 | 裂解 Cleavage |
| 21 | VIT_219s0014g02200 (VvNAC098) | 1.5 | gene=VIT_219s0014g02200 CDS=1105-2196 | 1826 | 1846 | 裂解 Cleavage | |
| 21 | VIT_219s0027g00230 (VvNAC021) | 1.8 | gene=VIT_219s0027g00230 CDS=317-1219 | 939 | 959 | 裂解 Cleavage | |
| 21 | VIT_214s0108g01070(VvNAC100-2) | 1.5 | gene=VIT_214s0108g01070 CDS=348-1439 | 988 | 1008 | 裂解 Cleavage | |
VvmiR164b (UGGAGAAGCAGGGCACAUGCU) | 21 | VIT_217s0000g06400(VvNAC100-1) | 1.0 | gene=VIT_217s0000g06400 CDS=732-1814 | 1375 | 1395 | 裂解 Cleavage |
| 21 | VIT_219s0014g02200 (VvNAC098) | 2.5 | gene=VIT_219s0014g02200 CDS=1105-2196 | 1826 | 1846 | 裂解 Cleavage | |
| 21 | VIT_219s0027g00230 (VvNAC021) | 1.8 | gene=VIT_219s0027g00230 CDS=317-1219 | 939 | 959 | 裂解 Cleavage | |
| 21 | VIT_214s0108g01070(VvNAC100-2) | 2.5 | gene=VIT_214s0108g01070 CDS=348-1439 | 988 | 1008 | 裂解 Cleavage |
| [1] |
|
| [2] |
doi: 10.3389/fpls.2017.00850 pmid: 28596775 |
| [3] |
崔梦杰, 王晨, 张文颖, 汤崴, 朱旭东, 李晓鹏, 房经贵. 无核葡萄研究进展. 植物生理学报, 2017, 53(3): 317-330.
|
|
doi: 10.1104/pp.53.2.317 |
|
| [4] |
|
| [5] |
|
| [6] |
doi: 10.1126/science.1159151 pmid: 18483398 |
| [7] |
doi: 10.1111/tpj.2017.92.issue-1 |
| [8] |
doi: 10.1242/dev.01206 pmid: 15226253 |
| [9] |
doi: 10.1093/jxb/ery172 |
| [10] |
doi: 10.1111/nph.v224.1 |
| [11] |
doi: 10.1242/dev.02817 pmid: 17287247 |
| [12] |
doi: 10.1105/tpc.106.045617 |
| [13] |
doi: 10.1093/jxb/eru072 |
| [14] |
doi: 10.1093/dnares/10.6.239 |
| [15] |
doi: 10.1105/tpc.9.6.841 |
| [16] |
|
| [17] |
doi: 10.1111/tpj.2005.44.issue-6 |
| [18] |
doi: 10.1007/s11105-008-0082-z |
| [19] |
|
| [20] |
doi: 10.1186/s12870-019-1719-9 |
| [21] |
吴伟民, 钱亚明, 赵密珍, 王壮伟, 袁骥. 赤霉素处理对‘魏可’葡萄果穗长度和坐果的影响. 江苏农业科学, 2006, 34(6): 257-258.
|
|
|
|
| [22] |
|
| [23] |
doi: 10.1371/journal.pone.0080044 |
| [24] |
doi: 10.1016/j.ydbio.2013.05.009 pmid: 23707900 |
| [25] |
doi: 10.1038/ng1791 |
| [26] |
doi: 10.3389/fpls.2018.00349 |
| [27] |
doi: 10.1007/s00438-018-1462-1 |
| [28] |
张文颖, 韩旭, 朱旭东, 解振强, 纠松涛, 黄雨晴, 贾海锋, 房经贵, 王晨. 葡萄miR159s靶基因的鉴定及其应答GA在果实不同组织的调控作用. 中国农业科学, 2019, 52(16): 2858-2870. doi: 10.3864/j.issn.0578-1752.2019.16.011.
doi: 10.3864/j.issn.0578-1752.2019.16.011 |
|
doi: 10.3864/j.issn.0578-1752.2019.16.011 |
|
| [29] |
宣旭娴, 盛子璐, 解振强, 黄雨晴, 巩培杰, 张川, 郑婷, 王晨, 房经贵. vvi-miR172s及其靶基因响应赤霉素调控葡萄果实发育的作用分析. 中国农业科学, 2021, 54(6): 1199-1217. doi: 10.3864/j.issn.0578-1752.2021.06.011.
doi: 10.3864/j.issn.0578-1752.2021.06.011 |
|
doi: 10.3864/j.issn.0578-1752.2021.06.011 |
|
| [30] |
王文然, 王晨, 解振强, 贾海锋, 汤崴, 崔梦杰, 房经贵. VvmiR397a及其靶基因VvLACs在葡萄果实发育中的作用分析. 园艺学报, 2018, 45(8): 1441-1455.
doi: 10.16420/j.issn.0513-353x.2018-0059 |
|
|
|
| [31] |
doi: 10.1093/nar/gkt1016 |
| [32] |
pmid: 27660497 |
| [33] |
doi: 10.1093/jxb/ert324 pmid: 24085581 |
| [34] |
doi: 10.1105/tpc.105.030841 |
| [1] | 李元晶, 袁瑞祥, 李永泰, 孙天歌, 刘峰, 李艳军, 张新宇. 棉花黄萎病菌β-葡萄糖苷酶基因的鉴定及其在致病中的功能[J]. 中国农业科学, 2026, 59(7): 1380-1399. |
| [2] | 张东梅, 周鑫鑫, 肖桂林, 曾祥国, 王春燕, 王泽先, 韩永超. 草莓花器应答灰霉病菌侵染的表现特征及抗病性评价方法[J]. 中国农业科学, 2026, 59(7): 1456-1466. |
| [3] | 朱嘉伟, 关璇, 饶博涵, 刘秀海, 范国元, 武运, 陶永胜. 短乳杆菌LB-21介导的酒精-苹乳三菌共发酵对干红葡萄酒色泽的影响[J]. 中国农业科学, 2026, 59(5): 1101-1110. |
| [4] | 马桂兰, 张旭阳, 李武. 鸟苷酸结合蛋白2在金黄色葡萄球菌诱导巨噬细胞凋亡中的调控作用[J]. 中国农业科学, 2026, 59(4): 912-926. |
| [5] | 张梦博, 谭鸿冰, 沈甜, 徐美隆, 周新明, 房玉林, 鞠延仑. 不同灌溉量与抗蒸腾剂处理对葡萄酒品质的影响[J]. 中国农业科学, 2026, 59(2): 413-426. |
| [6] | 冯伟晴, 倪媛蒨, 费腾, 李有梅, 谢兆森. 不同果形葡萄果实维管束形态结构、分布特征及其水分运输功能差异[J]. 中国农业科学, 2026, 59(1): 161-178. |
| [7] | 王思琪, 邹利人, 白瑞雯, 闫可, 王思洋, 齐晓光, 申海林, 温景辉. 赤霉素调控‘蜜汁’葡萄穗轴硬化关键基因的挖掘[J]. 中国农业科学, 2026, 59(1): 179-189. |
| [8] | 蒲丽霞, 张佳芮, 叶建萍, 黄秀兰, 樊高琼, 杨洪坤. 二氢赤霉素与秸秆覆盖对旱地小麦分蘖成穗与产量的影响[J]. 中国农业科学, 2025, 58(9): 1735-1748. |
| [9] | 谭西北, 兰徐颖, 刘崇怀, 樊秀彩, 姜建福, 孙磊, 李鹏, 余书鑫, 张颖. 不同抗性葡萄响应白腐病侵染的次生代谢物变化[J]. 中国农业科学, 2025, 58(9): 1767-1778. |
| [10] | 陈龙云, 胡俊强, 何灿, 史建荣, 徐剑宏, 王刚. 利用大孔吸附树脂与高速逆流色谱纯化脱氧雪腐镰刀菌烯醇-3-葡萄糖苷[J]. 中国农业科学, 2025, 58(8): 1627-1637. |
| [11] | 汤学燊, 党仕卓, 周娟, 李佳豪, 李梅花, 胡豪, 张亚红. 基于红蓝光调控分析VvBES1-1参与‘红地球’葡萄花芽的分化[J]. 中国农业科学, 2025, 58(8): 1650-1662. |
| [12] | 杨彩丽, 李永洲, 贺亮亮, 宋银花, 章鹏, 刘肇先, 李鹏慧, 刘三军. 葡萄TPS基因家族全基因组鉴定及VvTPS4在单萜形成中的功能验证[J]. 中国农业科学, 2025, 58(7): 1397-1417. |
| [13] | 张天雨, 李白, 藏金萍, 曹宏哲, 董金皋, 邢继红, 张康. 灰葡萄孢HMG家族基因的全基因组鉴定与表达规律分析[J]. 中国农业科学, 2025, 58(4): 704-718. |
| [14] | 郭奥琳, 林俊璇, 赖恭梯, 贺丽媛, 车建美, 潘若, 杨方学, 黄玉吉, 陈桂信, 赖呈纯. 过表达VdF3′5′H2对刺葡萄细胞花青素组分积累的影响[J]. 中国农业科学, 2025, 58(4): 802-818. |
| [15] | 从琪琪, 张静怡, 孟祥龙, 戴蓬博, 李波, 胡同乐, 王树桐, 曹克强, 王亚南. 我国苹果轮纹病弱毒菌株中病毒的鉴定及其携带情况的检测[J]. 中国农业科学, 2025, 58(3): 478-492. |
|
||